Microbial Identification based on Mass Spectral Libraries and Interspectral Distances
Introduction
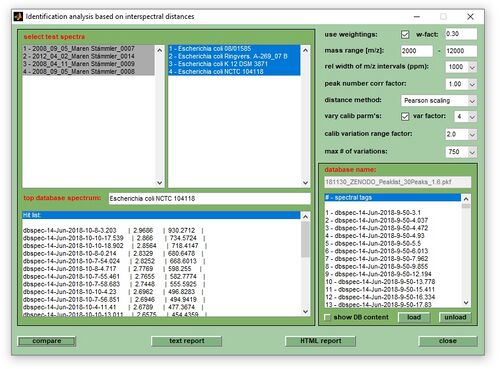
- In the MicrobeMS software package microbial identification based on mass spectral libraries and interspectral distances, or similarity measures such as D-values, is carried out by means of the subfunction cmpr (compare). The cmpr function is available from the Analysis menu bar (select compare with database), or by pressing the button compare with DB in the ANALYSIS tab (see Screenshot of MicrobeMS). The function calculates interspectral distances (on the basis of peak tables) between spectra in a database and the experimental mass spectra. Spectra in a database, also called reference spectra, are recorded from microorganisms with a known taxonomic status and can be of two types: (i) single experimental spectra and (ii) so-called database spectra. The latter are created from a selection of experimental spectra (see section interspectral distances below).
- Reference spectra are ranked by the cmpr function according to their spectral distances to the test spectrum such that the reference spectrum with the smallest distance (or highest score) appears on the top of the ranking list. As the genus, species and strain identity of the microorganisms used to record the database spectra are known, this allows microbial identification at certain taxonomic level.
- In MicrobeMS spectral distances can be obtained by using different metrices: Euclidean distances, D-values derived from Pearson’s product-momentum correlation coefficients, covariance and Pareto scaling (see section interspectral distances below). All distance algorithms search for matching peaks in the peak tables of the test spectra and the spectra comprising the database, respectively.
- The function cmpr always requires the presence of peak tables. These peak tables must be obtained from both types of spectra, test and reference spectra. Peak tables contain the m/z positions of the peaks, intensities of the unprocessed spectra, normalized intensities (also called weightings) and – in case of database spectra – frequency values of the mass peaks. Frequency values indicate how often a peak was found in the experimental spectra used to create the actual database spectrum.
- An important problem of calculating interspectral distances from different MALDI TOF mass spectra is to define the conditions at which a peak in the test spectrum matches a peak in a reference spectrum. It is known that MALDI TOF MS generally suffers from a relatively low precision of the experimentally determined m/z peak positions. Therefore, when analyzing spectra from biological and technical replicates mass peaks of theoretically identical m/z positions may be detected at slightly varying m/z positions. To deal with such inaccuracies of the m/z positions, a program wide variable ppm has been introduced. The ppm parameter defines the maximum allowed variation between theoretically identical peaks obtained by different measurement. The ppm variable defines the width of mass regions, or m/z sections (intervals), in which spectra are subdivided: [M-ppm/2 M+ppm/2], with M being the m/z position of a given section center. Only in cases where peaks from distinct spectral measurements fall within the borders of such m/z sections they are treated as identical peaks.
- Database spectra are ideally created from a defined number of experimental mass spectra, usually between 3 and 20. The procedure of creating database spectra always starts from raw experimental mass spectra and includes spectral pre-processing, peak detection followed by a statistical analysis of the peak tables. A database spectrum is essentially represented by a peak table in which the following values are stored:
-
a) the average peak position of mass peaks,
b) the mean intensities of the mass peaks (obtained from experimental mass spectra),
c) normalized peak intensities (weightings): the sum of the normalized intensities equals 100 in a database spectrum and
d) the frequency the peaks are found in the experimental mass spectra used to create the database spectrum.
(see also format of peak tables)
- Parameters for automated spectral pre-processing and peak detection are stored in the file 'microbems.opt'. This file is a simple text file which can be edited by text editors like Notepad. It is required to restart MicrobeMS to initialize changes made to this file. Note that existing blocks of pre-processed spectra, or peak tables are not overwritten when creating database spectra from experimental mass spectra.
Interspectral distances
In MicrobeMS interspectral distances are calculated from peak tables (see above) and can be of the following types:
- Euclidean distances: probably the most commonly chosen type of distance. The Euclidean distance can be considered the geometric distance in a multidimensional space, see Euclidean distance for details.
- Pearson: Pearson distances D1(x,y) between two peak tables x and y are calculated on the basis of Pearson's product momentum correlation coefficient, which is basically the covariance cov(x,y) of the two vectors divided by the product of their standard deviations (σx × σy, this product is also known as the total joint variance). Values of r1(x,y) vary between -1 (perfect negative linear correlation), 0 (no correlation) and 1 (perfect positive linear correlation). To obtain the Pearson distance D1(x,y) the following formula is applied: D1(x,y) = 1000 × (1-r1(x,y)). Pearson distance varies between 0 (identity - perfect positive linear correlation) and 2000 (anti-correlation - perfect negative linear correlation).
- Pareto (0.75): The Pareto-0.75 distance D¾(x,y) between two peak tables x and y is obtained on the basis of the Pareto-scaled correlation coefficient r¾(x,y) which is calculated by dividing the covariance cov(x,y) of the two vectors by the product of their standard deviations to the power of 0.75: r¾ = cov(x,y) / (σx × σy)^0.75. r¾(x,y) values are then scaled by divding them by r¾(x,x). For this purpose, the test peak table vector is compared with itself. D¾(x,x) can be subsequently computed from the following equation: D¾(x,y) = 1000 × [1-(r¾(x,y) / r¾(x,y))]. Note that the Pareto-0.75 distance can be smaller than 0 and larger than 2000.
- Pareto (0.50): The Pareto-0.50 distance D½(x,y) between two peak tables x and y is determined in a similar way to the Pareto-0.75 distance: The only difference is that the exponent value of 0.75 is replaced by a value of 0.50 (see above). Specifically, the Pareto-0.50 distance D½(x,y) is obtained on the basis of the Pareto-scaled correlation coefficient r½(x,y) which is calculated by dividing the covariance cov(x,y) of the two vectors by the product of their standard deviations to the power of 0.50: r½(x,y) = cov(x,y) / (σx × σy)^0.50. r½(x,y) values are then scaled by divding them by r½(x,x). For this purpose, the test peak table vector is compared with itself. D½(x,y) can be subsequently computed from the following equation: D½(x,y) = 1000 × [1-(r½(x,y)/ r½(x,x))]. The Pareto-0.50 distance can be smaller than 0 and larger than 2000.
- Pareto (0.25): The Pareto-0.25 distance D¼(x,y) between two peak tables x and y is determined similarly to the Pareto-0.75 distance: The only difference is that the exponent value of 0.75 is replaced by a value of 0.25 (see above). Like the Pareto-0.75 and the Pareto-0.50 distances, values for D¼(x,y) can be smaller than 0 and larger than 2000.
- Covariance: First, the covariance cov(x,y) between the two peak table vectors x and y is calculated. Then, the covariance cov(x,x) between the test peak table vector with itself is obtained. The covariance-based distance D0(x,y) is determined by the following equation: D0(x,y) = 1000 × [1-(cov(x,y) / cov(x,x))]. Covariance-based distances can be smaller than 0 and larger than 2000 .
Scores
Score values are directly computed from the inter-spectral distance values (D1, D¾, D½, D¼ and D0) by means of the following equation: Score = 1000 - D(i). Score values below one are set to one. In consequence score values may vary between 1 (negative, no, or almost no correlation), 1000 (identity), or values larger than 1000 (only in case of Pareto and covariance scaling, the checkbox use weigthings must be activated and w-fact is set to values larger than 0).
Computation of logarithmic score values log10(score) produces values between 0 (negative, no, or almost no correlation), 3 (identity), or above 3 (only in case of Pareto and covariance scaling). In MicrobeMS scores and log(scores) are used to assess and compare levels of similarity between the experimental mass spectra and microbial reference spectra in the database.
Note that the score values obtained by MicrobeMS should not be compared with the score values of Bruker's MALDI Biotyper: Due to the different algorithms used - MicrobeMS score values are based on interspectral distances - MicrobeMS scores tend to be larger than the corresponding MALDI Biotyper score values. Of note, such higher scores do not indicate better matches between test and database mass spectra.
Vary calibration parameters
Calibration parameters are varied when this checkbox is activated. When calculating distance values between test and reference spectra, the comparison is done for a set of [2n+1 × 2n+1] variations of three calibration constants, with n being the value chosen from the pulldown menu var factor (see screenshot at the top of this page). For example, if the default value of n=4 has been selected from the pulldown menu, MicrobeMS will calculate in each comparison interspectral distances between the respective reference spectrum and 729 test spectra [9 × 9 × 9 = (2 × 4+1) × (2 × 4+1) × (2 × 4+1)] representing the different combinations of three different calibration constants. In the reports, only one - the best (highest) - match will be displayed for each test spectrum.
Vary calibration range factor: this factor defines the range of variation in which the calibration factors are allowed to vary. High values indicate a large range and vice versa. Chose high values in case of badly calibrated spectra. Of note that a high calibration range factor may result in accidental high scores from non-related microbial taxa.
Max number of variations: This parameter is useful to reduce the computational load when calibration factors are varied. If the number of variations is larger than indicated a distance-based algorithm removes similar combinations of calibration factors.
Peak number corr factor (still experimental!): A factor for a still highly experimental procedure to compensate for different numbers of peaks derived from test and database spectra. Evidently, the ratio between peak numbers of test and database spectra has a strong impact on the distance values and therefore on the resulting scores. Select a factor of 1 (default) to deactivate this algorithm.
Use weightings
Allows to "weight" the influence of MS intensity values on the spectral distances, scores and thus, the identification results. If the checkbox use weightings is activated, interspectral distances can be obtained on the basis of intensity values. Otherwise, distances are calculated from barcode spectra. In cases where the checkbox use weightings is checked, the edit field w-fact (weighting factor) becomes active. Weigting factors may vary between 0 (barcode spectra) and 1 (full intensities) and define the relative weight of the MS intensity values when creating spectra for distance value calculation. For this the following formula is used: int(distspec) = (1-wfact)*mean(int(exp)) + wfact*int(exp) [wfact denotes the weighting factor, int(distspec) the intensity of a given peak in the spectrum for distance calculation, mean(int(exp)) the mean peak intensity of the original spectrum and int(exp) the original peak intensity].
Visualization and interpretation of the results
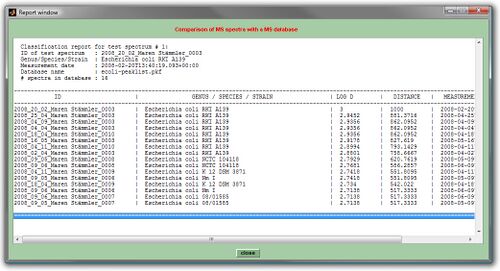
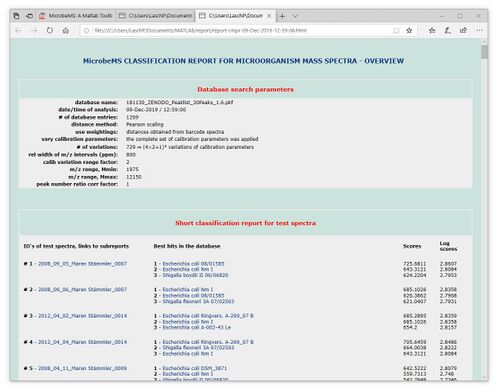
The results of the pattern matching analyses are provided as a score ranking list, either in a text, or a HTML format. In both types of lists, the top matching database entries are displayed on top position. Further records are listed below according to the scores achieved (see screenshots below).
While reports in the simple text format cannot be printed, HTML reports are printable for documentation purposes by using the appropriate function of the webbrowser software (Microsoft Internet Explorer, Mozilla Firefox, Opera, etc.). In cases where a pdf printer driver is available reports can be directly converted into a pdf format. Furthermore, all HTML reports are stored per default in a subfolder /report which is automatically created in the program's root directory (Windows). The name of the HTML report file will be of the format report-cmpr-DAY-MONTH-YEAR-HOUR-MIN-SEC.html, for example report-cmpr-19-Jun-2015-11-36-39.html.
Automatic workflow compare mass spectra against a spectral database
MicrobeMS allows identification of microorganisms based on MALDI-TOF mass spectra and mass spectral libraries by an automated and a manual workflow. This section describes the necessary procedures and steps required for automated identification.
- Load the mass spectral data files via the load spectra (Bruker data file format), [Import Mass Spectra in a mzXML Data Format|import spectra from mzXML data]], or the load MS multifile options of the File menu bar.
- For identification select the respective spectra in the listbox in the top left corner (the listbox is labeled by MicrobeMS spectra ID`s). To select multiple spectra hold the <shift> key while selecting.
- Start the automated identification procedure by pressing the button identification in the ANALYSIS tab (bottom of the main figure), or by choosing identification from the Analysis menu bar. The shortcut for this function is <Shift> + I. MicrobeMS performs then automated pre-processing and auto peak picking using the parameters defined in the configuration file of MicrobeMS, microbems.opt. Note that existing pre-processing data and peak tables are not overwritten by this function.
- When pre-processing / peak detection has been completed MicrobeMS will load the mass spectral database defined in microbems.opt and open a figure labeled as identification analysis based on interspectral distances (see the section Introduction at the top of this page. If the database cannot be loaded (e.g. because of wrong settings in microbems.opt) the programs offers to load this file manually.
- In the identification window modify the parameters and settings used for distance calculation then press compare (bottom, right). Press this button immediately to start the identification procedure with the default settings. Depending on the number of spectra and the size of the database the computation time may vary between a few seconds and several minutes. A progress indicator will be shown to give an idea of the work remaining. For a description of the parameters and settings see the section above.
- When classification has been finished the buttons text report and HTML report will be activated. Press either of them to see the reports (please refer to the section Visualization and Interpretation of the Results for more details).
Manual workflow compare mass spectra against a spectral database
In this chapter the manual workflow for identifying microorganisms based on their MALDI-TOF mass spectra and mass spectral libraries is described.
1. Load the mass spectral data files via the load spectra (Bruker data file format), import spectra from mzXML data, or the load MS multifile options of the File menu bar.
2. Manual spectral pre-processing: select first the respective spectra in the listbox in the top left corner (the listbox is labeled by MicrobeMS spectra ID`s). Hold the <shift> key to select multiple spectra while selecting. Spectral pre-processing can be started by pressing the appropriate buttons of the functions smooth (smoothing of spectra), baseline (baseline subtration), normalize (normalization), or calibrate (auto-calibration). Additional pre-processing procedures which can be applied to the spectra before peak picking are cut spectra and reduce resolution. Both functions are available from the Pre-processing menu bar. Recommended spectral pre-processing routines before peak detection are (Bruker spectra in the m/z range 2000 - 20,000): a) Smoothing with 21 smoothing points b) Baseline subtration (number of intervals: 60 - 100) c) Normalization (no parameters required) In selected cases additional pre-processing procedures may be useful.
3. Perform manual peak detection
4. Note that spectra selected for identification should contain valid peak tables. Spectra without associated peak table cannot be processed
5. Start the identification procedure by pressing the button compare with database in the ANALYSIS tab (bottom of the main figure), or by choosing compare with database from the Analysis menu bar. The shortcut for this function is <Shift> + H. MicrobeMS will then open a figure labeled as Identification analysis based on interspectral distances (see the section Introduction at the top of this page).
6. Load a mass spectral database by pressing the load button. After loading the content of the database can be printed in the command line window by pressing the button show DB content. Use unload to unload the database.
7. In the identification window modify the parameters and settings used for distance calculation then press compare (bottom, right). Press this button immediately to start the identification procedure with the default settings. Depending on the number of spectra and the size of the database the computation time may vary between a few seconds and several minutes. A progress indicator will be shown to give an idea of the work remaining. For a description of the parameters and settings see the section above.
8. When classification has been finished the buttons text report and HTML report will be activated. Press either of them to see the reports (please refer to the section Visualization and Interpretation of the Results for more details).