Identification Analysis by Means of LC-MS¹ and ''in silico'' Databases
Introduction
Microbial identification analysis by means of LC-MS¹ data and an in silico database represents an alternative bottom-up proteomics approach that involves (i) efficient extraction of proteins from cultivated microbial samples, (ii) digestion by trypsin and (iii) LC-MS measurements. Next steps in the analytical pipeline are (iv) extraction of MS¹ data followed by (v) systematic tests of MS¹ data against a library of strain-specific synthetic peptide mass profiles computed from UniProt Knowledgebase (UniProtKB) databases (i.e. Swiss-Prot and TrEMBL). These tests involves computation of score values derived from spectral distances between experimental and the in silico peptide mass data and compilation of score ranking lists. The taxonomic position of the microbial sample can be then determined by using the best-matching database entries.
The basic concept and main ideas of microorganisms identification analysis by LC-MS¹ and in silico databases are outlined in the following publication:
Identification of Microorganisms by Liquid Chromatography-Mass Spectrometry (LC-MS1) and in Silico Peptide Mass Libraries Lasch P, Schneider A, Blumenscheit C, Doellinger J Mol Cell Prot, 2020 19(12), 2125-2139
Requirements
- MS¹ spectral data obtained by LC-MS measurements from microbial extracts. An example of a LC-MS¹ test set (Burkholderia-LC-MS-set_03.muf) has been uploaded to Zenodo, see link below. The test set is stored in the spectral multifile (*.muf) format of MicrobeMS.
- The in silico database of strain specific peptide mass data (1.4 GB). This database can be downloaded from Zenodo, see link below. The current database version is stored in the *.pkf format and comprises more than 12,000 strain-specific profiles, each containing tens of thousands of peptide mass entries. Details of the *.pkf format are outlined here: Description of the MicrobeMS *.pkf file format
- MicrobeMS v. 0.82 (pcode), see Downloading MicrobeMS. Of note, Matlab pcode requires a valid Matlab license (R2014a or later, Windows or LINUX 64-bit). Furthermore, Matlab's Statistics Toolbox as well as the Bioinformatics Toolbox are required.
- Hardware requirements are substantial - more than 100 GB of RAM are highly recommended when testing the in silico database by experimental data!!
Download link:
In silico Database for Identification of Microorganisms by Liquid Chromatography-Mass Spectrometry (LC-MS¹) Lasch P, Schneider A, Blumenscheit C, Doellinger J, Zenodo (December 13, 2019) doi: 10.5281/zenodo.3573996 https://doi.org/10.5281/zenodo.3573996
Performing identification analysis
This chapter describes the workflow for identifying microorganisms based on their LC-MS¹ tryptic peptide patterns and a in silico library. The workflow follows the ideas outlined here: Microbial identification by means of mass spectral libraries and interspectral distances. For testing the identification workflow it is recommended to use the LC-MS¹ test data set Burkholderia-LC-MS-set_03.muf and the in silico database available from Zenodo (non-redundant-proteomes-22000-processed-Trembl+Swissprot.7z, 7zip archive). See the next section to learn how to prepare own LC-MS¹ data for identification.
- Start the pcode version of MicrobeMS v.082 or later by typing 'mass' at the Matlab command prompt. Analysis of LC-MS¹ data requires special software settings. Please activate first the LC-MS mode of MicrobeMS by chosing settings → LC-MS settings from the File menu bar.
- Load the LC-MS¹ data file (*.muf format) via the load MS multifile option of the File menu bar. Note that *.muf files should contain a valid peptide peak table (field spec.pik) while spectral data stored in the fields spec.org and spec.pre are ignored. When LC-MS¹ data are imported a command line message will be given (LC-MS spectra can not be plotted). For further details, refer to spectral multifile (*.muf) format of MicrobeMS
- Do NOT pre-process the data
- Do NOT perform peak detection
- Select the respective LC-MS¹ spectra for identification from the listbox displayed in the top right corner of the MicrobeMS gui.
- Start the identification procedure by pressing the button compare with database in the ANALYSIS tab (bottom of the main figure), or by choosing compare with database from the Analysis menu bar. The shortcut for this function is <Shift> + H. MicrobeMS will then open a figure labeled as Identification analysis based on interspectral distances.
- Load the in silico database (*.pkf format) by pressing the load button. After successful loading the content of the database can be printed in the command line window by pressing the button show DB content. Use unload to unload the database.
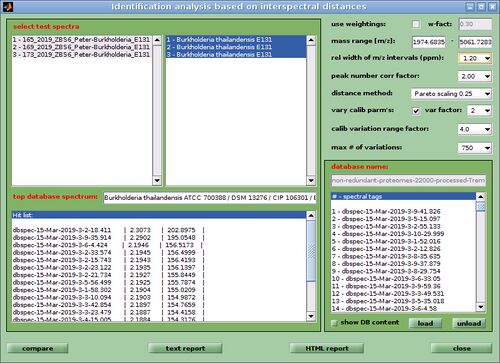
- In the identification window modify the parameters and settings used for distance calculation. Recommend parameters are shown in the screenshot of the window identification analysis based on interspectral distances (see right). Press the button compare to start the identification procedure. Depending on the number of spectra, the size of the database and the type of computer hardware the computation time may vary between 90 seconds and several minutes. A progress indicator will be shown to give an idea of the work remaining. For a description of the parameters and settings see the section Microbial identification by means of mass spectral libraries and interspectral distances.
- When classification analysis has been finished the buttons text report and HTML report will be activated. Press either of them to see the reports. Please refer also to the section Visualization and Interpretation of Identification Results. Example of a HTML result file which is obtained when using files and settings of the example used: https://wiki.microbe-ms.com/uploads/report-cmpr-20-Dec-2019-18-07-16.html
Preparing own LC-MS¹ test data
Identification analysis by MicrobeMS and an in silico library of microbial tryptic peptide mass data requires LC-MS¹ test data in a special data format. In the following it is outlined how to prepare the experimental data.
Prerequisites
1. Tab-separated text files containing the experimental LC-MS¹ data
2. Matlab R2014a or newer
3. readlcmstxtfile.m - a Matlab function for data format conversion
Preparing tab-separated text files
Peptide feature detection of the MS¹ data can be carried out using the Minora algorithm of the Proteome Discoverer software from Thermo-Fisher. Peptide feature text files should contain, among others, the following parameters: MS¹ peak positions (in m/z units) with the respective ion charge state. Two examples of such peptide feature text files can be downloaded here:
https://wiki.microbe-ms.com/uploads/165_2019_ZBS6_Peter-Burkholderia_E131_ConsensusFeatures.zip
https://wiki.microbe-ms.com/uploads/169_2019_ZBS6_Peter-Burkholderia_E131_ConsensusFeatures.zip
Download readlcmstxtfile.m
https://wiki.microbe-ms.com/uploads/readlcmstxtfile.m
Using the function readlcmstxtfile.m
- Start Matlab R2014a or newer
- Navigate to the directory where readlcmstxtfile.m resides and type
- As a result a standard file open dialog box for choosing the source text file comes up. Select the respective LC-MS¹ peak file and press open.
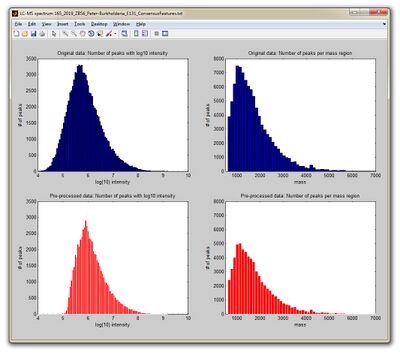
- The Matlab function readlcmstxtfile.m creates now a new window entitled as LC-MS spectrum + name_of_the_text_file showing four different plots, see screenshot to the left. Furthermore, a variable pik is returned which contains LC-MS¹ peak positions and intensities.
- In the next step it is now required to append the array pik to an existing *.muf file. This can be achieved by loading a *.muf file of choice in Matlab, for example by typing
>> pik = readlcmstxtfile(ithreshperc);
at the Matlab command prompt. ithreshperc is a scalar between 0 and 150 with an impact on the number of MS¹ peaks to be removed: the higher the value of ithreshperc, the more peaks are removed, i.e. the lower the number of signals. If unsure use values between 50 and 70 as a start.
Furthermore, the function readlcmstxtfile can be also used to find and remove peaks due to aa side chain modifications (oxidation, deamidation) and allows selection of ion charge states (default: +2 - +4). For more details refer to the m-code of the function readlcmstxtfile.m.
- Add then the array pik to the structure array spec by typing
>> load('Burkholderia-LC-MS-set_03.muf','-mat')
at Matlab's command prompt. This will create a new structure array spec; the file Burkholderia-LC-MS-set_03.muf can be downloaded from Zenodo, see link above.
- Store the *.muf spectra multifile file by typing
>> spec(1).pik = pik;
Note that this will overwrite the existing pik table of the first entry of the structure array spec. You want to modify then also other fields of spec(1). Existing spectral data, for example in the fields spec(1).org or spec(1).pre are ignored. Please refer also to the description of the *.muf file format.
>> save('Burkholderia-LC-MS-set_XX.muf','-mat','-v7.3');
at the Matlab command prompt. The *.muf file can be used in the same way as outlined in the example given in the previous section (Performing identification analysis).